SECTION 1: MULTIPLE CHOICE QUESTIONS (With All Options)
1. What roles does nicotinamide adenine dinucleotide play in oxidative pathways?
(a) Enzyme
(b) Coenzyme ✅
(c) Prosthetic group
(d) Inhibitor
Correct Answer: (b) Coenzyme
Explanation: NAD⁺ acts as a coenzyme that transfers electrons in oxidation reactions.
2. The enzymes that catalyse reactions in which two molecules are joined together by synthesis of new bonds using energy from ATP are placed in group:
(a) Hydrolase
(b) Ligase ✅
(c) Lyase
(d) Transferase
Correct Answer: (b) Ligase
Explanation: Ligases join molecules and use ATP energy.
3. Which of the following is an example of hydrolases?
(a) Lipase ✅
(b) Glycogen phosphorylase
(c) Pyruvate decarboxylase
(d) Cytochrome oxidase
Correct Answer: (a) Lipase
Explanation: Lipase breaks fats by addition of water.
4. Which of the following statements about enzymes is correct?
(a) They increase the activation energy of a reaction
(b) They are consumed during the reaction
(c) They are specific in terms of the reactions they catalyse ✅
(d) They always work optimally at high temperatures
Correct Answer: (c) They are specific in terms of reactions they catalyse
5. Enzyme B requires Zn²⁺ to catalyse conversion of substrate X. Zinc is best identified as a:
(a) Coenzyme
(b) Activator ✅
(c) Substrate
(d) Product
Correct Answer: (b) Activator
Explanation: Metal ions like Zn²⁺ act as activators (inorganic cofactors).
6. If an enzyme solution is saturated with substrate, the most effective way to obtain an even faster yield of products would be:
(a) Add more enzymes ✅
(b) Add more substrate
(c) Add an allosteric inhibitor
(d) Add a non-competitive inhibitor
Correct Answer: (a) Add more enzymes
7. How does a non-competitive inhibitor decrease the rate of an enzyme-catalysed reaction?
(a) By binding the active site of the enzyme
(b) By changing the shape of the enzyme ✅
(c) By changing the free energy change of the reaction
(d) By acting as a coenzyme for the reaction
Correct Answer: (b) By changing the shape of the enzyme
8. Which enzyme class is responsible for catalysing the addition of water to a substrate molecule?
(a) Ligase
(b) Lyase
(c) Hydrolase ✅
(d) Isomerase
Correct Answer: (c) Hydrolase
SECTION 2: SHORT QUESTIONS
- Define enzyme and co-factor. Enzyme: A biological catalyst (mostly protein) that lowers activation energy of specific reactions without being consumed. Co-factor: Non-protein helper (ion or organic molecule) required by some enzymes for activity. Apoenzyme + cofactor = Holoenzyme.
- Differentiate between co-enzyme and prosthetic group. Co-enzyme: Loosely bound, easily separated, act as carriers (NAD, CoA). Prosthetic group: Tightly/permanently bound part of enzyme (Heme, FAD).
- Hydrolases: Class 3 enzymes that catalyse hydrolysis (break bonds by adding water). Examples: Lipase (fats), Amylase (starch).
- Activation energy: Minimum energy needed for reactants to form transition state and start reaction.
- Feedback inhibition: End product of a pathway inhibits the first enzyme (usually allosterically) to prevent overproduction.
- Competitive inhibitors: Malonic acid, Sulphonamides. Non-competitive: Cyanide, Hg²⁺, Pb²⁺.
- Optimum pH: pH at which enzyme shows maximum activity. Pepsin: 2.0, Salivary amylase: 6.8, Trypsin: 8.0.
SECTION 3: LONG QUESTIONS
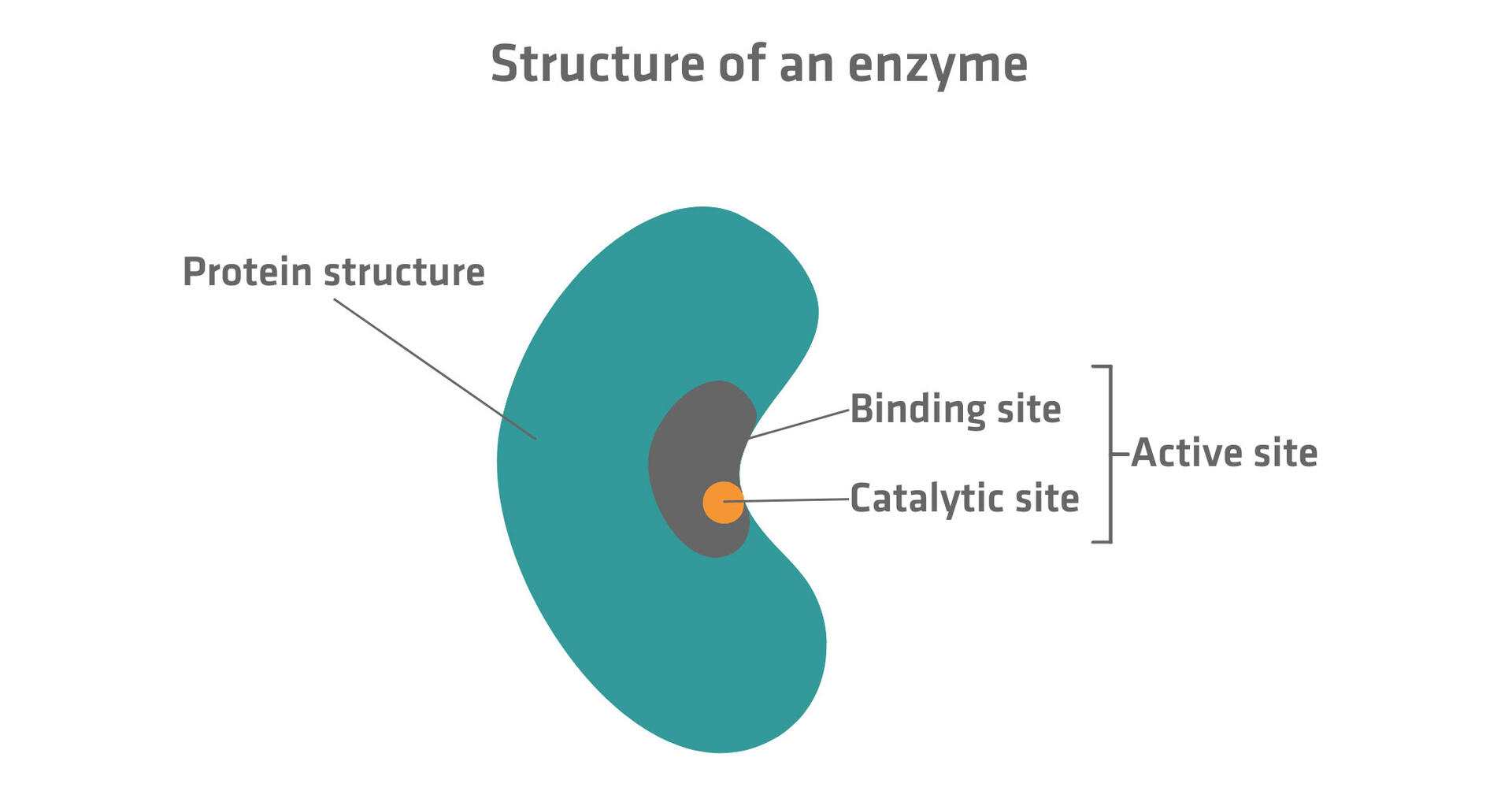
Describe the structure of enzyme, explaining the role and component parts of the active site of an enzyme. Enzymes are mostly globular proteins with precise 3D shape. They have apoenzyme (protein) + cofactor. Active site is a small pocket/cleft formed by few amino acids.
- Binding site: Holds substrate by weak bonds (specificity).
- Catalytic site: Amino acids that actually break/make bonds. Role: Specificity, lowers Ea, orients substrate perfectly.
Diagram: Structure of Enzyme & Active Site
Differentiate among the three types of co-factors, by giving examples. (i) Inorganic ions (activators): Mg²⁺, Zn²⁺, Fe²⁺. (ii) Co-enzymes: Loosely bound organic (NAD, CoA). (iii) Prosthetic groups: Tightly bound organic (Heme, Biotin).
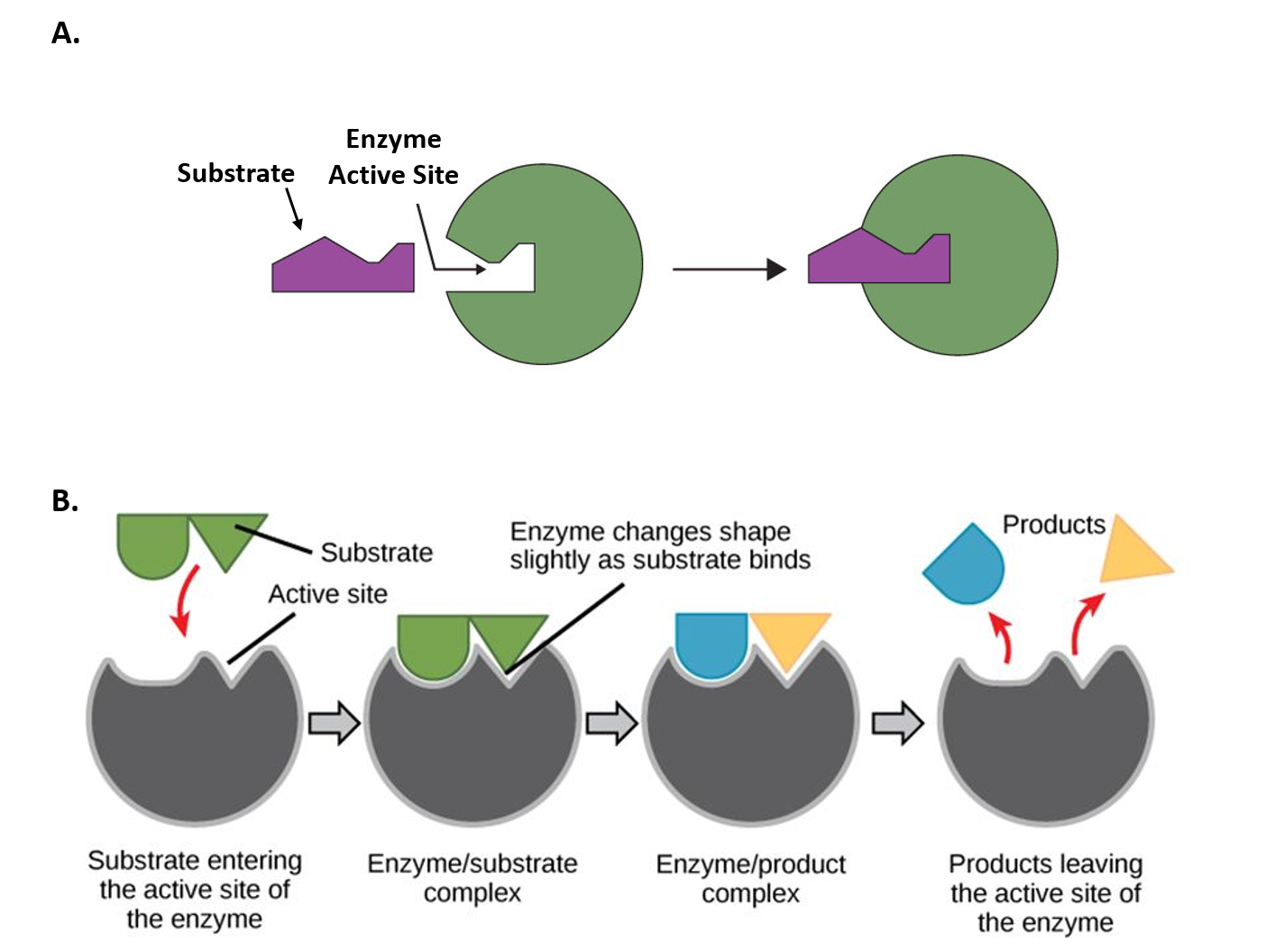
Explain the mechanism of enzyme action through Induced Fit Model, comparing it with Lock and Key Model. Lock & Key (Fischer 1894): Rigid active site, exact fit like key in lock. Induced Fit (Koshland 1958): Enzyme is flexible. Substrate binding causes conformational change so active site fits perfectly (like hand in glove). Better explains catalysis and regulation.
Comparison Diagram (Lock & Key vs Induced Fit)
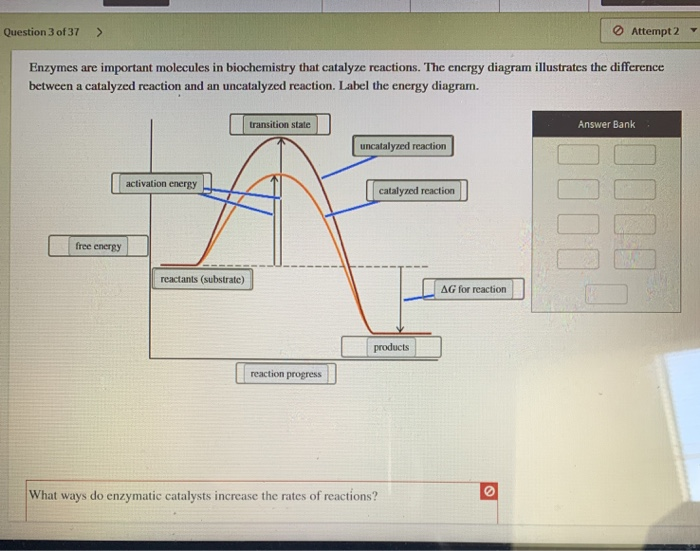
Define activation energy and explain through graph how an enzyme speeds up a reaction by lowering activation energy. Activation energy = minimum energy barrier for reaction to start. Enzyme lowers Ea by stabilising transition state → more molecules react at body temperature. ΔG remains same.
Activation Energy Graph
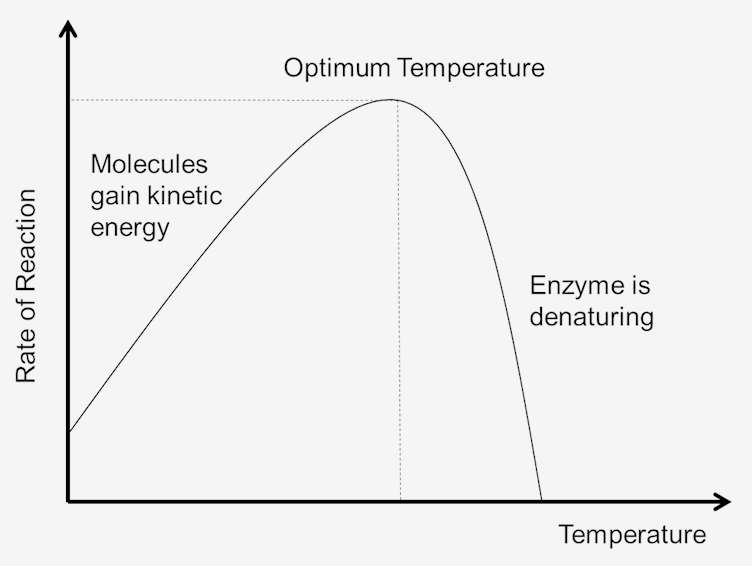
Describe the effect of temperature on the rate of enzyme action. Rate ↑ with temperature up to optimum (more kinetic energy, more collisions). At optimum (37°C humans) = max activity. Above optimum → denaturation (H-bonds break), rate falls sharply.
Temperature Effect Graph (Bell curve)
Compare the optimum temperatures of enzymes of human and thermophilic bacteria. Human: ~37°C (denature >45°C). Thermophilic bacteria: 60–80°C or more (stable due to extra bonds). Example: Taq polymerase (72°C) used in PCR.
Describe how the concentration of enzyme affects the rate of enzyme action. When substrate excess, rate directly proportional to [enzyme] (more active sites). Graph = straight line upward.
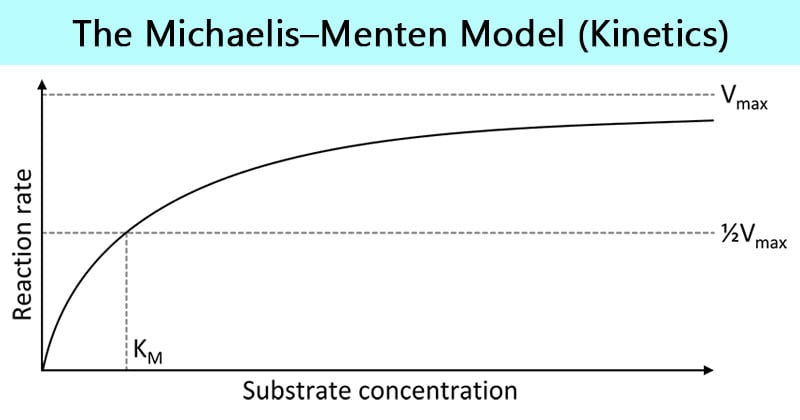
Explain the effect of substrate concentration on the rate of enzyme action. Low [S] → rate ↑ linearly. High [S] → reaches Vmax (all sites saturated). Km = [S] at ½ Vmax (affinity measure).
Michaelis-Menten Graph
9-10. Describe enzymatic inhibition, its types… Categorize inhibitors into competitive and non-competitive. Competitive: Bind active site, ↑ Km, Vmax same, overcome by more substrate. Non-competitive: Bind allosteric site, ↓ Vmax, Km same, cannot be overcome.
Inhibition Graphs
- Explain feedback inhibition. End product binds allosteric site of first enzyme → pathway stops. Saves energy. Example: Isoleucine inhibits threonine deaminase.
Feedback Inhibition Diagram
- Classify enzymes on the basis of the reactions catalysed. 6 classes (IUBMB): Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases.
- Give examples of enzymes’ naming according to substrates. Urease, Sucrase, Lipase, Amylase, Protease






.jpg)


+by+Blacklotus+Landscaping.jpg)



0 Comments